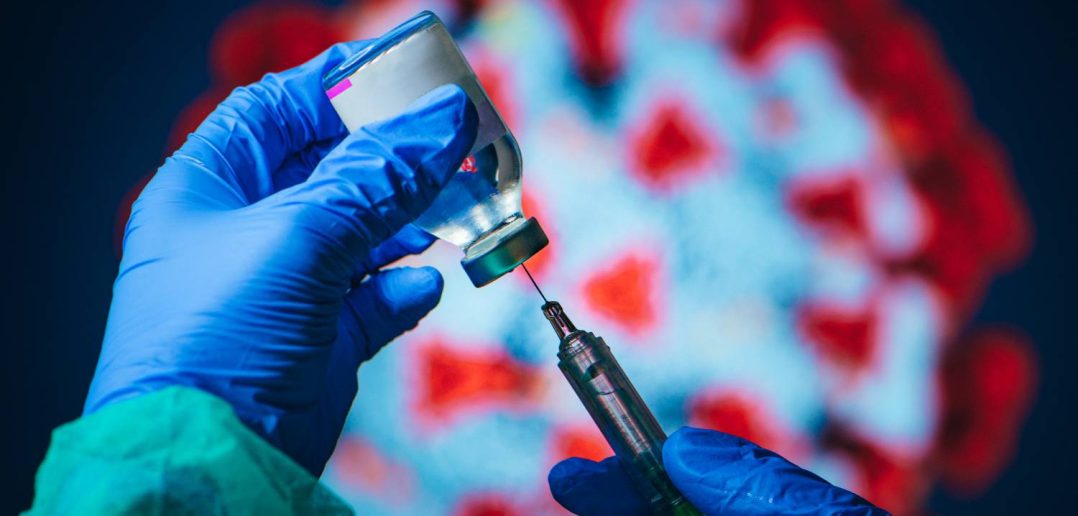
The pandemic caused public health agencies and life sciences companies to scramble to find effective treatments. Exploring the efficacy of existing drugs to treat COVID, an example of what's known as drug repurposing, was an intense focus for Qais Hatim, a Data Scientist at the US Food and Drug Administration (FDA).

I spoke with Hatim about his research, its applications to COVID and beyond, and his use of machine learning and data visualization to help nontechnical experts understand possible new applications of proven drugs.
What is the focus of your work at FDA?
Qais Hatim: I have a couple roles. As a liaison between industry and the FDA, I try to understand current problems within FDA and what industry, or other federal agencies, are doing to solve the problems. I try to bring these two sectors together, keeping in mind the privacy of FDA data and confidentiality of the information, of course.
I also work as a lead data scientist, so I develop projects, manage them and look for opportunities to apply techniques from simple statistical modeling to advanced deep learning and machine learning analysis. I help nontechnical people within the FDA expedite their use of current innovations. For instance, I may use nontechnical subject matter experts to advance the technical problem so that there is science and technology working together.
I understand you’ve done some work around what is called computational drug repurposing. Can you explain what that is?
Hatim: This is a very recent work we’ve done with SAS. In general, computational drug repurposing is a drug discovery approach that uses computational methods to identify new uses of existing drugs.
There are three main advantages in drug repurposing:
- To identify a new target for existing drugs which can save time and money in the drug development processes. Clinical trials can take years to develop, but using drug repurposing you can really cut the time and money for developing new drugs.
- Also, it can be used to predict the efficacy of existing drugs for new indications, which can help to reduce the risk of failure in clinical trials. Sometimes you will be successful in your clinical trials, and sometimes you will spend a lot of money but cannot get to the target you are looking for.
- Finally – and this is really important – drug repurposing can identify new drug combinations. Sometimes you need more than one drug to treat certain diseases and this process helps us understand drug interactions and drug combinations for treating disease.
There are several different computational methods used for drug repurposing. One common approach which we use SAS for is use machine learning to identify patterns in the data that can be used to predict the efficacy of drugs for a new indication.
Another approach, which is common in industry, is to use in silico screening. In silico screening is a biological experiment conducted on a computer or via computer simulation that uses virtual tools to make predictions about the behavior of different components in order to identify potential drug targets for diseases.
As the world scrambled to figure out treatments for COVID-19, that had to be a game changer for drug repurposing. Can you tell us how COVID-19 provided unique opportunities to implement computational drug repurposing approaches?
Hatim: It was my honor to work with an emergency task force at the White House during the pandemic. COVID-19 created a sense of urgency in the scientific community and led to the rapid development of new computational methods for drug repurposing.
We used different methods to identify potential drug candidates for COVID-19 much more quickly than if we waited for traditional drug discovery approaches. In addition, and this was really important, as we were working in this fast-paced emergency task force, we had a large data set of data on COVID-19 itself. This was very helpful for the training and validation of the computational drug repurposing models. We had data on the virus itself as well as data about the human body’s response to the virus.
Several drugs that were identified through computational drug repurposing are now used for treating COVID-19. These drugs include remdesivir, baricitinib and molnupiravir.
COVID-19 really opened our eyes to how to use computational models in identifying the drugs that exist to treat this pandemic. We also built a community that is ready to encounter future pandemics using these models for discovering drugs. But hopefully, we will never need them.
How did the work during COVID support advancements in the treatment of rare diseases, and particularly for populations that don't typically enroll in clinical trials?
Hatim: Drug repurposing can support advancement in the treatment of rare disease and of populations that don't typically enroll in clinical trials in a number of ways.
Rare diseases are often difficult to treat because there are small markets for drugs that target them, which makes it difficult for pharmaceutical companies to invest in development of new treatments. Drug repurposing can actually help to overcome this challenge by identify existing the drugs that may be effective.
In addition, drug repurposing can help to improve the safety and efficacy of treatments for rare diseases. Rare diseases are often difficult to study because there are few patients with the diseases. So this makes it difficult to conduct clinical trials to assess the safety and efficacy of treatments. Drug repurposing can help overcome this challenge by identifying existing drugs with a long history of safety and that have been studied in large populations.
Related: Read more stories from SAS bloggers on COVID-19
Drug repurposing also can help to make treatment for rare diseases more affordable. Clinical trials are not cheap. To develop of a new treatment is a very cost costly process and the resulting drug will likely be unaffordable for a patient with a rare disease. With drug repurposing we can model what already exists on the market to identify existing drugs that have already been approved for other applications and may be effective with a rare disease.
Finally, we can increase the diversity of patients who participate in a clinical trials. Historically speaking, most clinical trials are conducted on a specific population: white males of European descent. Other populations may be ignored, which can introduce bias into the clinical trials. Drug repurposing can help increase the diversity of patients who participate in clinical trials by identifying existing drugs that have been studied on more diverse populations.
How could this project help improve public health and support FDA's mission?
Hatim: FDA and the National Institutes of Health (NIH) are both actively involved in drug repurposing. For example, the Office of Orphan Products Development, or OOPD, and the Office of Regenerative Medicine (ORM) are both funding projects around drug repurposing.
The OOPD, for example, is funding research using drug repurposing for new treatments of rare diseases. The Office of Regenerative Medicine is funding research on new treatments for diseases that are caused by defects in the body’s ability to repair or regenerate tissues.
NIH actually has a repurposing drug discovery program, or RDDP, that is funding research on new uses for existing drugs.
They’ve successfully funded a number of projects that resulted in promising new treatments for malaria, HIV and cancer.
Another example in the FDA is a drug named sirolimus that was originally developed to prevent organ rejection in transplant patients. Now this is also approved for the treatment of rare genetic disorder called tuberous sclerosis complex. Also, NIH is investigating the use of metformin, a diabetes drug, to treat Alzheimer's diseases.
There is a very big scope in using drug repurposing because it's cheaper and there is a lot of data. So you can really investigate a new innovation or new drugs for treating some diseases that you’ve never used a particular drug for.
In your drug repurposing research, what did you think you would find? Can you tell us a little bit about your hypothesis?
Hatim: So working with SAS and the subject matter experts in the FDA, our central hypothesis is that we can find new candidate drugs to treat diseases by searching for similar patterns between existing drugs that have been used to treat a given disease or similar diseases.
We started with a data set from Drug Bank, from which we can retrieve information about relationships between drugs and proteins. We used the technology to build a graphical network. This is the essence of the work: a graphical network for drugs and proteins.
We also used different public data sources but no private data. We really needed to show that you can use what’s available in public data to develop the model. Though this could also be a proof of concept model that can be implemented with private data.
We also used data on biochemical entities from a database named STRING, which gives us a relationship between different proteins.
All total, we used five databases, so the problem became logistical. How can we put them in one database so that we can perform network analysis? We have a large XML database with thousands of attributes per drug.
We work with the data engineering team from SAS and experts from FDA to extract the attributes that help our analysis. Extracting the attributes was not solely based on technology. There was a human in the loop who brought expertise to the trials.
We were able to link data on drugs, genes and diseases in the database and present them visually. The great thing about our work is that we give nontechnical experts a visual way to see similarities between drugs, proteins, genes and virus information to target one drug or another.
Our ability to simplify the problem was an important part of the hypothesis, in addition to what we thought we could accomplish technically with SAS.
You talked a little bit about your work with SAS. Can you dive a little deeper into that? How has SAS helped in your research both on the technology side but also just support from SAS personnel?
Hatim: There are several projects I’ve developed with SAS since joining the FDA in 2015. For instance, we’ve done projects on drug-induced liver injury, deep learning, text mining for patient narrative to extract adverse event and analyze possible root causes drug repurposing, of course, and drug labeling that we’ve submitted to SAS Explore.
The SAS team supports us not only from the technical side but also step-by-step in making this really successful. In this project we used SAS® Studio (and) SAS Visual Data Mining and Machine Learning on SAS® Viya®. This enabled us to deploy SAS in our runtime environment for data management and analytics. SAS Viya was very easy to implement and gave us the power and the scalability necessary for key analytical procedures for different techniques.
I also value the flexibility of SAS in connecting with open source. For example, we utilize Python in most of the work in data engineering. But Viya is not siloed; it has the capability to use open source alongside SAS.
We also leveraged graphical interfaces through SAS Visual Analytics to give access and information to non-technical experts.
I’m really grateful to the subject matter experts such as Professor Dr. Faraj M. Abdullah, Dean of Al-Manara College of Medical Science, whose support was critical to achieving this success.
I’d also want to thank the people from NIH who brought incredible subject matter expertise. Also, SAS R&D who quickly solved problems if they arose. We also had experts from the FDA Office of New Drugs which has really helped in making this a successful story.