A common task in statistics is to approximate a large data matrix by using a low-rank approximation. Low-rank approximations are used for reducing the dimension of a problem (for example, in principal component analysis), for image analysis via the singular value decomposition (SVD), and for decomposing large matrices that are encountered in text analytics. Mathematically, these problems are similar: you start with a data matrix containing hundreds or thousands of columns (variables in the data), and you reduce it to a handful of fundamental features.
This article looks at a two-factor approximation, where an n x m data matrix, Y, is approximated by the product L*R, where L is an n x k matrix and R is a k x m matrix. (As a mnemonic, I use L as the "left" matrix and R as the "right" matrix.) Usually k is a small integer. This is motivated by a particular two-factor decomposition, called the nonnegative matrix factorization (NMF), which I will write about in a future article. Although the SVD and the NMF share similarities, they are also very different. For most data matrices, you can standardize the factors in the SVD to obtain a unique decomposition. However, two-factor NMF decomposition is more complicated. This article takes a closer look at the nonuniqueness of the factors in a two-factor decomposition.
The nonuniqueness of the NMF explains why different algorithms might give NMFs that are correct but look completely different. The tips in this article make it easier to compare factorizations that are produced by different software or different algorithms.
The Fundamental Theorem of Linear Algebra
Nonuniqueness of factors is very common in mathematics. For example, the integer 12 can be expressed in multiple ways as the product of two nontrivial factors: 2*6, and 3*4. A way to deal with nonuniqueness is to standardize the factors so that they have special properties. For example, the Fundamental Theorem of Arithmetic says that every composite integer can be represented uniquely as a product of prime numbers, up to the order of the factors. This allows us to represent 12 = 22*3 in a standard way. Note that the representation has two properties: the factors are "standardized" to be prime, and the prime factors are listed in increasing order (2 before 3) to eliminate the order ambiguity.
In a similar way, the SVD decomposition of a matrix is sometimes called the Fundamental Theorem of Linear Algebra. The singular value decomposition states that every n x p matrix can be written as the product of three matrices: A = U Σ VT where U and V are orthogonal matrices, and Σ is a diagonal matrix that contains the singular values. The SVD is not unique, but if the singular values of Σ are distinct, then we can obtain a factorization that is "mostly unique," in the following sense:
- Order the singular values of Σ in decreasing order. You can always do this by using a permutation that re-orders the columns of U and V.
- The columns of U and V are unit length, but they can vary in sign. Make some arbitrary choice such as "the first row of U is positive" or "the sum of the columns of U are positive." Rules like these remove the sign ambiguity.
All statistical libraries order the singular values. However, most do not standardize the signs of the columns of U and V, which means that you might see slightly different SVDs if you compare the results from different software. However, if you apply a rule to standardize the columns of U, then the results from different software will agree, provided that the singular values are distinct.
The SVD has a strong connection to low-rank approximations. Namely, if you choose some integer k ≥ 1, you can use the SVD to find the best rank-k approximation to the original matrix, Y. The new decomposition is often denoted as UkΣkV`k, where Uk is defined as the first k columns of U, Σk is the upper-left k x k submatrix of Σ, and Vk is the first k columns of V.
Nonuniqueness of a two-factor decomposition: The scaling ambiguity
As explained above, the SVD provides a three-factor rank-k decomposition that can be standardized so that it becomes unique.
However, if you want to express the SVD by using only two factors, you must sacrifice uniqueness.
The SVD represents the linear transformation as the composition of a rotation, a scaling, and another rotation. You can try to distribute the scaling transformation (Σ)
into one or both of the orthogonal matrices, but there are infinitely many ways to do that.
This is called the scaling ambiguity.
If α is any value in [0, 1], you can form a two-factor decomposition in infinitely many ways:
Y = (U*Σα) (Σ1-αV`)
You can easily construct examples to demonstrate this nonuniqueness of a two-factor decomposition, as shown in the following SAS IML program:
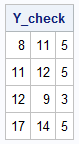
proc iml; reset fuzz; /* Define a simple matrix with an underlying rank of 2 */ Y = { 8 11 5, 11 12 5, 12 9 3, 17 14 5 }; /* Use SVD to find the factors */ call svd(U, Sigma, V, Y); /* Extract the first k=2 columns to get a rank-2 product */ i = 1:2; U2 = U[,i]; S = Sigma[i]; V2 = V[,i]; /* You can distribute the scaling of S into the left factor, the right factor, or both factors by varying alpha */ alpha = {0, 0.5, 1}; do j = 1 to nrow(alpha); L = U2 * diag(S##alpha[j]); R = T( V2 * diag(S##(1-alpha[j])) ); /* The product L * R is identical for all choices of alpha! */ prod = L*R; print prod; end; |
I do not display the output from the program, but for every assignment inside the DO loop, the product L*R is exactly the same and is equal to Y (because Y is rank-2).
A problem with two-factor decompositions
In general, suppose you decompose a matrix Y as Y = L*R, where L is n x k and R is k x m.
This decomposition is not unique because
you can also write
Y = (L*A) (A-1R)
for any invertible matrix, A.
Thus, let's examine ways to standardize the problem to resolve as many ambiguities as we can.
The order ambiguity
A permutation matrix
is an invertible matrix that changes the orders of columns or rows. The inverse of a permutation matrix equals its transpose.
Thus, for any two-factor decomposition Y = L*R, you can permute the columns of L and simultaneously permute the rows of R
by using a permutation matrix, P, as follows:
Y = (L*P) (P`*R).
This is known as the order ambiguity or the permutation ambiguity.
For example, the following IML program interchanges the order of the columns of L and the rows of R, and the product is the same:
/* Permute the columns of L and rows of R */ P = {0 1, 1 0}; L_new = L * P; R_new = P` * R; Y_check = L_new * R_new; /* perfectly reconstructs Y */ print Y_check; |
The mixing ambiguity
Recall that the SVD decomposition is (practically) unique is because the left and right factors are orthonormal. If you change the scaling of the U or V matrices, or you change the order of their columns and rows, they are still orthogonal. Consequently, if we adopt certain conventions (for example, sort the columns by the magnitude of the singular values and require that V is orthonormal), then we can standardize the SVD, provided that the singular values are distinct.
If we do not insist that the left and right factors be orthogonal, then we might have to sacrifice uniqueness of the factorization. As mentioned above, for any invertible matrix, A, it is true that Y = (L*A) (A-1R) is a valid two-factor decomposition. The following IML program shows an example:
/* A can be ANY invertible matrix. */ A = { 0.9 0.1, 0.1 0.9}; L_new = L * A; R_new = inv(A) * R; Y_check = L_new * R_new; /* the product is preserved */ print Y_check; |

Why is this important?
As mentioned earlier, a popular algorithm in machine learning and text mining is the nonnegative matrix factorization (NMF) algorithm. Given an n x m data matrix, Y, that has nonnegative elements, you can find factors L (an n x k matrix) and R (a k x m matrix) that also have nonnegative elements, and for which L*R is the best rank-k approximation to Y subject to the constraints that L and R are nonnegative. The constraints on L and R might make us hopeful that we cannot arbitrarily form new factors (L*A) and (inv(A)*R). That is often correct: Not all invertible matrices A will result in the new factors being nonnegative!
You can prove that if A is the product of a permutation matrix and a positive diagonal matrix, then the new factors are nonnegative, but, unfortunately, there are other choices of A that also preserve nonnegativity of factors.
For example, the following rank-2 example defines L and R as nonnegative factors. The A matrix is the same as in the previous section. The inverse of A is NOT nonnegative, but the products (L*A) and (inv(A)*R) retain their nonnegative properties. Consequently, the NMF factorizations are both correct. Furthermore, you cannot transform one into the other simply by scaling and sorting the columns of L or the rows of R.
L1 = {1 2 , 3 8 , 5 4 }; R1 = {22 21 20 19, 12 22 11 10}; Y = L1 * R1; print Y, L1, R1; A = {0.9 0.1, 0.1 0.9}; invA = inv(A); print A, invA; /* Note: inv(A) is not nonnegative! */ /* nevertheless, the products (L*A) and (inv(A)*R) can still be nonnegative */ L2 = L1*A; R2 = inv(A)*R1; Y_check = L2 * R2; print Y_check, L2, R2; |
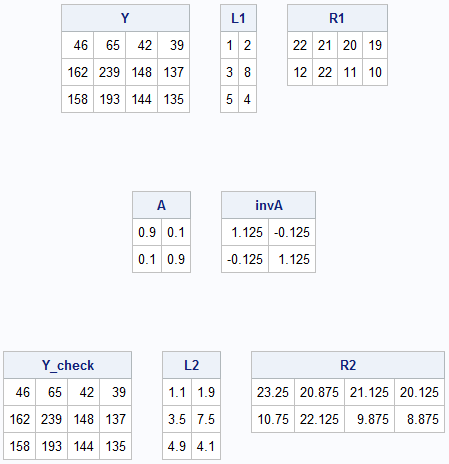
This example motivates an important question: is there is a way to check whether two decompositions (L1, R1) and (L2, R2) represent the same solution up to multiplication by an invertible matrix? The answer is yes! If all the matrices are full rank and L1*R1 = L2*R2, then you can solve a least-squares system to find the matrix A that converts one solution into the other.
Here is how to find the matrix A.
Assume that there is a k x k matrix A for which
L2 = L1 * A
R2 = inv(A) * R1
You can use
the generalized inverse (also called the Moore-Penrose inverse)
to solve for A in each equation.
In the first equation, multiply on the left by
the Moore-Penrose pseudoinverse for L1.
In the second equation, multiply on the right by
the Moore-Penrose pseudoinverse for R1.
If you get the same answer from each equation, then the solutions are equivalent.
/* What if we use different algorithms and one algorithm returns (L1,R1) whereas the other returns (L2,R2)? Can we find an invertible matrix, A, that maps one solution to the other? */ A_from_L = ginv(L1) * L2; A_from_R = R1 * ginv(R2); /* Do the two LS solutions give the same matrix? */ max_diff = max(abs(A_from_L - A_from_R)); print max_diff; print A_from_L, A_from_R; /* Note: inv(A) is not nonnegative! */ |
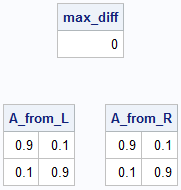
In this example, each overdetermined system produces the same matrix, so the two factorizations are equivalent up to a linear transformation.
Summary
This article explores the uniqueness of various matrix factorizations related to rank-k approximation. If you use the classical SVD algorithm, you can obtain a rank-k approximation of a data matrix that is mostly unique. To get uniqueness, you must specify how to resolve the order ambiguity and the scaling ambiguity. If a matrix has distinct singular values, you can obtain a unique rank-k decomposition that you can use to compare SVDs from different software. The factors in the SVD are orthogonal matrices, which imposes enough restrictions on the factors that you can obtain a unique representation.
Unfortunately, uniqueness is not guaranteed for other decompositions. An example is the NMF, which I will discuss in a future post. In the NMF, the factors are not orthogonal, but they are composed of nonnegative elements. You can resolve the scaling ambiguity and the order ambiguity by standardizing the factors. However, two algorithms might give different factors whose product is the same. In that case, you can solve a least squares system to determine whether there is an invertible linear transformation that convert one set of factors into the other set.
