When there are two equivalent ways to do something, I advocate choosing the one that is simpler and more efficient. Sometimes, I encounter a SAS program that simulates random numbers in a way that is neither simple nor efficient. This article demonstrates two improvements that you can make to your SAS code if you are simulating binary variables or categorical variables.
Simulate a random binary variable
The following DATA step simulates N random binary (0 or 1) values for the X variable. The probability of generating a 1 is p=0.6 and the probability of generating a 0 is 0.4. Can you think of ways that this program can be simplified and improved?
%let N = 8; %let seed = 54321; data Binary1(drop=p); call streaminit(&seed); p = 0.6; do i = 1 to &N; u = rand("Uniform"); if (u < p) then x=1; else x=0; output; end; run; proc print data=Binary1 noobs; run; |
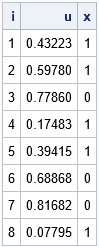
The goal is to generate a 1 or a 0 for X. To accomplish this, the program generates a random uniform variate, u, which is in the interval (0, 1). If u < p, it assigns the value 1, otherwise it assigns the value 0.
Although the program is mathematically correct, the program can be simplified. It is not necessary for this program to generate and store the u variable. Yes, you can use the DROP statement to prevent the variable from appearing in the output data set, but a simpler way is to use the Bernoulli distribution to generate X directly. The Bernoulli(p) distribution generates a 1 with probability p and generates a 0 with probability 1-p. Thus, the following DATA step is equivalent to the first, but is both simpler and more efficient. Both programs generate the same random binary values.
data Binary2(drop=p); call streaminit(&seed); p = 0.6; do i = 1 to &N; x = rand("Bernoulli", p); /* Bern(p) returns 1 with probability p */ output; end; run; proc print data=Binary2 noobs; run; |
Simulate a random categorical variable
The following DATA step simulates N random categorical values for the X variable. If p = {0.1, 0.1, 0.2, 0.1, 0.3, 0.2} is a vector of probabilities, then the probability of generating the value i is p[i]. For example, the probability of generating a 3 is 0.2. Again, the program generates a random uniform variate and uses the cumulative probabilities ({0.1, 0.2, 0.4, 0.5, 0.8, 1}) as cutpoints to determine what value to assign to X, based on the value of u. (This is called the inverse CDF method.)
/* p = {0.1, 0.1, 0.2, 0.1, 0.3, 0.2} */ /* Use the cumulative probability as cutpoints for assigning values to X */ %let c1 = 0.1; %let c2 = 0.2; %let c3 = 0.4; %let c4 = 0.5; %let c5 = 0.8; %let c6 = 1; data Categorical1; call streaminit(&seed); do i = 1 to &N; u = rand("Uniform"); if (u <=&c1) then y=1; else if (u <=&c2) then y=2; else if (u <=&c3) then y=3; else if (u <=&c4) then y=4; else if (u <=&c5) then y=5; else y=6; output; end; run; proc print data=Categorical1 noobs; run; |
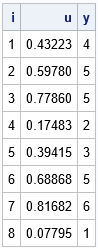
If you want to generate more than six categories, this indirect method becomes untenable. Suppose you want to generate a categorical variable that has 100 categories. Do you really want to use a super-long IF-THEN/ELSE statement to assign the values of X based on some uniform variate, u? Of course not! Just as you can use the Bernoulli distribution to directly generate a random variable that has two levels, you can use the Table distribution to directly generate a random variable that has k levels, as follows:
data Categorical2; call streaminit(&seed); array p[6] _temporary_ (0.1, 0.1, 0.2, 0.1, 0.3, 0.2); do i = 1 to &N; x = rand("Table", of p[*]); /* Table(p) returns i with probability p[i] */ output; end; proc print data=Categorical2 noobs; run; |
Summary
In summary, this article shows two tips for simulating discrete random variables:
- Use the Bernoulli distribution to generate random binary variates.
- Use the Table distribution to generate random categorical variates.
These distributions enable you to directly generate categorical values based on supplied probabilities. They are more efficient than the oft-used method of assigning values based on a uniform random variate.

2 Comments
Thanks for pionting to the table distribution again. I am using that sicne one of your earlier posts and it is quite nice way to handle this type of distribution. However, I have experienced that quite a number of runs per simulation are needed to stabilize the outcome around the expected probabilities. Any way to increase that efficiency as well?
Thanks for writing. The variance of the estimates depends on several factors. Some you can control and some you can't:
1. The sample size. The parameter estimates in large samples tend to be closer than the estimates in small samples. Said differently, the standard error of an estimate decreases when the sample size increases.
2. The number of Monter Carlo simulations. For some statistical tests and procedures, you can get good estimates by using a moderate number of simulations. For example, you can probably use 1,000 MC simulations to get good estimates for a univariate mean whereas you might need 100,000 to get good estimates for kurtosis. That is because the variance of the mean estimator is much smaller than the variance of the kurtosis estimator.
So there is not a simple solution to your problem. Some statistics require many MC simulations, others do not.