Locating missing values is important in statistical data analysis. I've previously written about how to count the number of missing values for each variable in a data set. In Base SAS, I showed how to use the MEANS or FREQ procedures to count missing values. In the SAS/IML language, I demonstrated the COUNTN and COUNTMISS functions that were introduced in SAS/IML 9.22.
But did you know that you can also use the COUNTN and COUNTMISS functions to determine the number of missing values in each observation? This can be useful in an analysis (such as regression) in which you need to delete observations that contain one or more missing value. Or it can be useful to determine observations that contain missing values for a large number of variables.
To begin, let's count missing values in the SAS DATA step by using the CMISS function.
Count missing values in the DATA step
For the SAS DATA step, there is a SAS Knowledge Base article that shows how to use the CMISS function to count missing values in observations. The following example uses an array of variables and the CMISS function to count the numbers of missing values in each observation:
/* define 6 x 3 data set; rows 2, 4, and 5 contain missing values */ data Missing; input A B C; datalines; 2 1 1 4 . . 1 3 1 . 6 1 . 1 . 3 4 2 ; data A; set Missing; array vars(3) A--C; /* contiguous vars */ numMissing = cmiss(of vars[*]); run; proc print; run; |
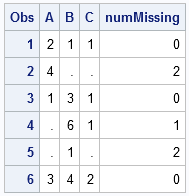
It is the ARRAY statement that makes the CMISS function convenient. If the variables are contiguous in the data set (as in this example), you can use the double-dash notation (A -- C) to specify the variables. If your variables share a common prefix, you can use the colon wildcard character (:) to specify the variables. For other useful techniques to specify an array of variable names, see the SUGI 30 paper, "Techniques for Effectively Selecting Groups of Variables" by Stuart Pollack.
Count missing values in each row in the SAS/IML language
In one of my first blog posts, I showed how to use the SAS/IML language to remove observations with missing values. However, that post was written prior to the release of SAS/IML 9.22, so now there is an easier way that uses the COUNTMISS function. The COUNTMISS function has an optional second parameter that determines whether the function returns the total number of missing values, the number in each column, or the number in each row. The following statements define a matrix with missing values and count the number of missing values in each row:
proc iml; use Missing; read all var _NUM_ into x; close Missing; rowMiss = countmiss(x, "ROW"); /* returns 0,1,2 or 3 for each row */ print rowmiss; |
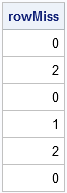
The output shows that the returned value is a vector with the same number of rows as x. The vector contains the number of missing values in each row of x.
It is now easy to use the LOC function (another function that I've written about often) to find the rows that contain missing values:
/* which rows have one or more missing values? */ jdx = loc(rowMiss>0); print jdx; |

Similarly, it is easy to extract the subset of rows that contain no missing values:
idx = loc(rowMiss=0); x = x[idx,]; /* delete rows that contain a missing value */ |
The x data matrix is the listwise deletion of the original data. You can now use the x matrix in statistical data analysis in PROC IML.

5 Comments
Another [albeit less-efficient] way to count missing obs by row, is to hard code missing-data indicators and sum. I prefer this way because you are all set to go when you want to understand whether the structure of missing-ness is dependent on a set of covariates (to test MAR/MCAR/..) or when constructing an inverse-weighted missing-data probability regression, for example.
Useful to know about these count functions - thanks. The IML example reminds me of a situation where I have been burnt before. Above I would have coded loc(rowMiss) rather than loc(rowMiss>0), it makes no difference in your example (apart from a little faster run time!). But once I had a bug in a program that was due to coding loc(x^=0), eventually I realised that the x^=0 part is not only redundant, but it also throws away the masking of missing values that is built into plain loc(x).
Thanks for the tip about missing values! I write the comparison operators in my LOC calls so that the code is easier to read, but the correct handling of missing values is a nice bonus. These fast O(n) operations won't make any difference in the performance of an O(n^2) or O(n^3) algorithm.
For numeric variables, will this count all values that are <= .Z ?
What about for character variables? will it count if _char_ in ('', ' ', '.')?
Pingback: Visualize missing data in SAS - The DO Loop