Suppose that you have a SAS/IML matrix and you want to set each element of a submatrix to zero (or any other value). There is a simple syntax that accomplishes this task.
If you subscript a matrix and do not specify a row, it means "use all rows." So, for example, you can assign zeros to the third column of a matrix by using the following syntax:
proc iml; x = j(3, 5, 1); /* create 3 x 5 matrix of ones */ x[ , 3] = 0; /* assign zeros to third column (all rows) */ print x; |
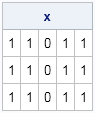
In the same way, if you do not specify a column, it means "use all columns." So, for example, you can assign zeros to the second row of a matrix by using the following syntax:
x[2, ] = 0; /* assign zeros to second row (all columns) */ |
Of course, you can extend these examples by assigning nonconstant vectors to rows or columns:
x[3, ] = 1:5; /* assign {1 2 3 4 5} to third row */ |
But here's a cool tip that many SAS/IML programmers do not know: you can fill the entire matrix by skipping the rows and the columns! For example, to assign zeros to entire matrix (regardless of dimensions), use the following syntax:
x[ , ] = 0; /* assign zero to all elements */ print x; |

I don't see this syntax used very often. It is never used on the right-hand side of an assignment because the expression t = x is easier and clearer than t = x[ , ]. However, when assigning values into a pre-allocated matrix, it is an efficient alternative to x = j(nrow(x), ncol(x), 0) or x[1:nrow(x), 1:ncol(x)] = 0. The "double empty" technique also preserves any matrix attributes that you might have assigned by using the MATTRIB statement, whereas using the J function deletes the old matrix and creates a new one.
Do you have a favorite SAS/IML syntax that is tricky or seldom-used? Share it in a comment!

8 Comments
Rick, I have been experimenting with setting all the elements of larger matrices to a constant and I can't understand why x=j(100, 1, 0) * j(1, 30, 0) is around 4 times faster than j(100, 30, 0), and incidentally a little faster than the empty subscripts method. How can making one 3000 element array and filling with zero take longer than the creation of 3130 elements in 3 arrays plus the overhead of all the multiplications?
I do not know. Let me think about this.
Dear Dr.Rick, I need your help. I already bought your 1st iml book and already ordered your next book. I know It has everything I need. But it will reach to me in end of next month, too late for me. I need to generate a data set for given correlation structure. I got a macro (I think it is also written by you) programme,gives from NORMAL dist. I need use this for different distribution like cauchy, chisquared,lognormal,web,....Please can you tell me how to do this.
I regret that I cannot provide personal assistance through this blog. I recommend that you post your question to a SAS Support Community at https://communities.sas.com/community/support-communities
I understood..Thank you Rick.
OK, I thought about it. I think what you were seeing was a consequence of multithreading of matrix multiplication (including outer products z=x`*y). When SAS first started multithreading SAS/IML, we started with matrix multiplication. This result makes sense if your CPU has four cores. I think when you reported this issue in 2013, you were using a version of SAS/IML for which matrix multiplication was running on 4 cores, but the J function was still single threaded. I just tried your problem using SAS/IML 14.1 (for which the J function is multithreaded) and the J function is now faster, as you would expect.
Rick, I need to create a matrix having 734 rows and 17 columns the row and column total are fixed how do i create such matrix having random values obeying the restriction i mentioned...Please help
You can ask programming questions like this at the SAS Support Communities. For this question, use the J function to allocate a matrix with 734 rows and 17 columns. Then use the RANDGEN subroutine and specify the distribution (uniform, normal, exponential,...) from which you want to draw the random values.